9:40am
This page describe the techniques our transmission electron microscopes can provide. Visit Scanning electron microscopy techniques for SEM techniques.
Conventional TEM
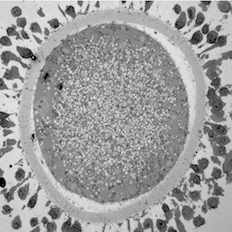
Maturing sheep oocyte by Karen Reader, read more below.
A beam of electrons is transmitted through an ultra thin specimen, interacting with the specimen as it passes through. The transmission of unscattered electrons is a function of the specimen thickness and elemental composition. Areas of the specimen that are more electron dense allow fewer transmitted unscattered electrons and appear darker, conversely the thinner areas and those containing lighter elements, permit more transmission and appear lighter. From this we can learn about the morphology of the specimen such as the size, shape and arrangement of the particles which make up the sample as well as their relationship to each other.
TEM can also provide crystallographic information about the sample such as the arrangement of atoms in the specimen and their degree of order, including detection of atomic-scale defects in areas a few nanometres in diameter. To prepare samples for TEM the specimens are typically fixed in glutaraldehyde, stained with uranyl acetate, embedded in resin and then cut into sections 70-80nm thin. Depending on the sample, the CM100 TEM is able to resolve images down to a few nanometres.
Image of maturing sheep oocyte by Karen Reader
I have been studying the ultrastructural differences between adult and prepubertal lamb oocytes to find reasons for the reduced ability of juvenile oocytes to produce viable embryos. This oocyte is surrounded by expanding cumulus cells and a zona pellucida layer. The cytoplasm is full of mitochondria, lipid droplets, transparent vesicles, and an MII nucleus.
Correlative TEM
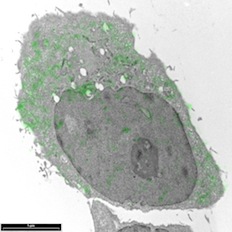
Breast cancer cells by Sharon Lequeuex, read more below.
Correlative microscopy involves using multiple microscope systems to observe the same specimen, most commonly light and transmission electron microscopy. This technique pairs the high resolution of the electron microscope which is able to reveal the cellular ultrastructure with the ability to follow specific targets on or in living cells, revealing dynamic localisation and/or function of target molecules using fluorescence microscopy.
Ultrathin sections of tissue are treated with a label that is able to be imaged under both microscopes, e.g. fluorescence photo-oxidation, quantum dots, double labelling with both fluorescent and gold conjugated antibodies, or fluoronanogold. The samples are first imaged using fluorescence microscopy and then transferred to the transmission electron microscope. Similar results can be achieved by taking parallel sections and preparing them either for light microscopy or transmission electron microscopy.
Other microscope systems can also be correlated, such as confocal and scanning electron microscopy (see array tomography), or scanning electron microscopy and transmission electron microscopy.
Image of breast cancer cells by Sharon Lequeux
Breast cancer cells that have been immunolabelled for YB1 protein with a green fluorescent dye conjugated to gold particles (we used Life Technologies Alexa Fluor® 488 Goat Anti-Rabbit IgG, 5 nm Colloidal Gold Conjugate). The cell can be looked at an even higher magnification to see which cellular structures the 5nm gold has tagged to.
Cryo-TEM
Using the cryo-TEM means samples can be viewed while frozen, avoiding chemical fixation and reducing damage to the samples caused by the electron beam.
Cryoultramicrotomy
Rather than being dehydrated and embedded in resin, tissue is lightly fixed and then suspended in a matrix (gelatin). It is then infiltrated with a cryoprotectant (sucrose) and frozen in liquid nitrogen. Frozen sections are then cut from the block onto grids. This preparation method is particularly good for immunolabelling.
Electron Tomography
By imaging of sections up to 300nm thick while tilting the specimen to 80° the tilt series can then be used to create a 3D reconstruction of the sample's internal structures.
Freeze Fracture TEM
 The freeze-fracture technique consists of freezing and then physically breaking apart (fracturing) a biological sample and visualising the exposed structural detail of the fracture plane by vacuum-deposition of platinum-carbon to make a replica for examination in the transmission electron microscope. Freeze fracture can provide three-dimensional perspectives of cellular organisation and details of membrane structure at macromolecular resolution. Of particular importance is the technique's ability to reveal the distribution and organisation of integral membrane proteins as intramembrane particles in the membrane plane.
The freeze-fracture technique consists of freezing and then physically breaking apart (fracturing) a biological sample and visualising the exposed structural detail of the fracture plane by vacuum-deposition of platinum-carbon to make a replica for examination in the transmission electron microscope. Freeze fracture can provide three-dimensional perspectives of cellular organisation and details of membrane structure at macromolecular resolution. Of particular importance is the technique's ability to reveal the distribution and organisation of integral membrane proteins as intramembrane particles in the membrane plane.
High Pressure Freezing (HPF)
Most biological specimens are highly hydrated (contain a lot of water) and this can be stabilised by freezing it to ice. Normal freezing is accompanied by the formation of ice crystals however, and these crystals destroy the fine structure of specimens. If the specimen is compressed to about 2000 bar at the moment of freezing, these crystals are suppressed and vitreous ice can be created. This preserves the water in the specimen without the damage caused by ice crystals. Further processing of the specimen (usually freeze substitution) can then proceed at low temperatures without the damage that results from processing at room temperature.
Immunocytochemistry
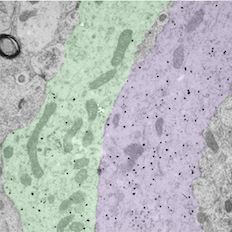
Bundled GnRH neuron dendrites by Rebecca Campbell, read more below.
An antigen is localised by one or more antibodies that are applied to the cells. An electron-dense tag is attached directly or (more commonly) indirectly to the last of the added antibodies. Some tags are organic molecules that possess structured electron opacity (e.g. ferritin), others are enzymes whose reaction product can be detected after the addition of the substrate (e.g. horseradish peroxidase), and yet others are heavy metals that can visualised directly (e.g. colloidal gold).
The only major requirement for localisation is that a specific antibody has been developed against the antigen. Other things to take into consideration are the size of the tag (whether it will infiltrate into the cellular structures), fixation and its effect on the antigenicity of the tissue, and whether to label before or after embedding. Pre-embedding labelling is best for surface labelling of live cells as antibodies do not penetrate cell membranes. Live cells may be broken open with detergents to reveal antigenic determinants (however ultrastructural preservation is compromised). It is also possible to use frozen ultrathin sections for immunocytochemistry. In the post-embedding technique cells are labelled on sections of tissue embedded in resin. Antigenicity may be compromised by fixation, but it is possible to etch tissue sections to expose antigenic sites.
Image of bundled GnRH neuron dendrites by Associate Professor Rebecca Campbell
The small black dots label the dendritic arms of two GnRH neurons (pseudocolored green and purple), cells in the brain that control fertility. The discovery that these two distinct dendritic elements, cut in the longitudinal plane, are found in close apposition and possess an attachment plate (shaded zone) between them, is suggestive of how these scattered cells communicate with one another.
Metal Shadowing

Myosin molecules by Richard Easingwood, read more below.
A thin layer of evaporated metal, such as platinum, is laid at an angle on a biological sample. An acid bath dissolves the biological material, leaving a metal replica of its surface, which can then be examined in the transmission electron microscope. Variations in the angle and thickness of the deposited metal allow an image to be formed because some incident electrons will be scattered in various directions rather than pass through the preparation. If the metal is deposited mainly on one side of the sample, for instance, the image seems to have "shadows" where the metal appears dark and the shadows appear light. This techniques creates a three dimensional view of the specimen and is used to obtain information about the shapes of purified viruses, fibres, enzymes, and other subcellular particles.
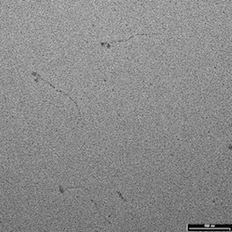
Image of myosin molecules by Richard Easingwood (specimen provided by John Harris)
Metal shadowing is a well established method for visualising small, isolated structures, such as macromolecules, in the TEM. The object here was to check a preparation for the correct configuration of myosin molecules. Replicas of purified myosin molecules were prepared for viewing by jet deposition on a molecularly-flat substrate. The molecules were then rotary shadowed at a low angle (6º) with platinum.
Negative Staining
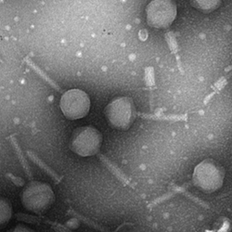
Isolated bacteriophage by Jin Lee.
In negative staining the macromolecule itself is usually unstained but is instead surrounded by a dense stain. As a result, the specimen appears in negative contrast (lighter in tone against a dark background). Negative stains are not used on sectioned materials, but are used to contrast whole, intact biological structures (viruses, bacteria, cellular organelles) that have been deposited on a supporting plastic or carbon film.
Image of isolated bacteriophage by Jin Lee
Microbiology undergraduate students isolate and purify a bacteriophage and then characterize it. This includes preparing the specimen for imaging in the TEM.
Scanning Transmission Electron Microscopy (STEM)
STEM (scanning transmission electron microscopy) enables the use of other of signals that cannot be spatially correlated in TEM, including secondary electrons, scattered beam electrons, characteristic X-rays, and electron energy loss—allowing very accurate elemental mapping and chemical analysis of samples.
Single Particle Reconstruction
Single particle reconstruction takes many images of viruses or proteins at different orientations and averages them together to produce a 3D reconstruction of the particle. The combined image has much stronger and easily interpretable features than individual images which contain much more noise. The nominal resolution of the new microscope is 0.27 nanometres.
