Image analysis overview
When analysing images, it's likely you'll have to make a subjective call on at least some of the steps in the process.
Typical questions are:
- Where should I decide the boundary is?
- Which intensity range should I choose to select my region of interest?
These are important questions to answer well, as they have a strong influence on your results.
It can be useful to think of every image as a collection of numbers. This may help you to proceed with caution. If you see them merely as images, it's easy to become entirely subjective on whether they are acceptable or not.
The programs listed below are a good starting point for many projects. If you are unsure of what to do, or need help in developing an analytical method, please feel free to make contact for advice.
Email confocal.microscopy@otago.ac.nz
What are the benefits of open-source software?
- You can be trained and then use the same software on your own computer
- Online support forums are often more comprehensive and supportive than commercial software
- They are often responsive to user requests for new or improved features
- File formats are not proprietary and therefore unlikely to become unreadable if the company collapses
- The program and updates are free
What about programs like Photoshop?
They are designed to make fabulous looking images – which they do very well. They are not designed to enable analytical objectivity to analyse your images. We recommend only using them for tasks such as cropping or tidying up rough edges.
3D Slicer
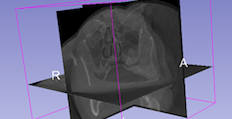 Designed for the medical profession, but 3D Slicer is perfectly capable with any other type of 3D data such as confocal, µCT (micro-computed tomography), and electron microscopy tomography files. In this respect it's similar to Fiji (below), but offers a different range of strengths, and weaknesses. Be prepared for a fairly steep learning curve.
Designed for the medical profession, but 3D Slicer is perfectly capable with any other type of 3D data such as confocal, µCT (micro-computed tomography), and electron microscopy tomography files. In this respect it's similar to Fiji (below), but offers a different range of strengths, and weaknesses. Be prepared for a fairly steep learning curve.
3D Slicer runs on any operating system and is free – but please acknowledge the software if you use it.
Getting started with 3D slicer
- Download 3D Slicer from their website
- Documentation resources (Slicer website)
This document outlines aligning pre- and post-operative 3D CT data sets and will help familiarise you with some of the basic functions of Slicer:
Semi-automatic alignment (PDF)
Amira
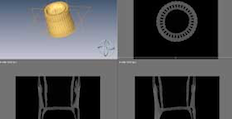 Amira is not open source but we have licenses. It can be very good for volume and surface rendering. It does have to be used on our designated systems.
Amira is not open source but we have licenses. It can be very good for volume and surface rendering. It does have to be used on our designated systems.
Blender
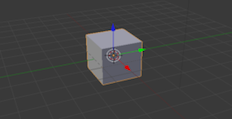 Blender is an open source program which has great capability for free software. Used primarily for 3D rendering and animations. If you're thinking of using it, plan well ahead to allow time to become familiar with it. The learning curve is very steep but the reason it's difficult is its capability with very complex tasks. Blender has a large and well informed community for support and inspiration.
Blender is an open source program which has great capability for free software. Used primarily for 3D rendering and animations. If you're thinking of using it, plan well ahead to allow time to become familiar with it. The learning curve is very steep but the reason it's difficult is its capability with very complex tasks. Blender has a large and well informed community for support and inspiration.
One note of caution with Blender is that it is designed to create spectacular images which can create a challenge when you are aiming to present your data with objectivity and precision.
It runs on MacOS, Linux, and Windows.
Download Blender from their website
Fiji / ImageJ
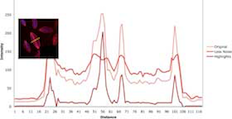 Fiji / ImageJ is a very underrated program, it can be expanded by installing any of the vast array of plugins and macros, which like the program itself are free as open source software. ImageJ's interface will perform many complex tasks and is platform independent.
Fiji / ImageJ is a very underrated program, it can be expanded by installing any of the vast array of plugins and macros, which like the program itself are free as open source software. ImageJ's interface will perform many complex tasks and is platform independent.
It's well worth checking the ImageJ plugins page for tools that may suit your needs.
An alternative, and functionally identical program is Fiji, which comes ready packaged with many plugins and will search for updates and install them for you.
The type of things typically done with Fiji / ImageJ include:
- Intensity and density levels
- Length, area, volume measurement
- Particle analysis
- Path tracing
- Volume rendering
Getting started
This PDF covers the basics of using ImageJ, how to open files, calibrate images, threshold, and where to find the program. Used as part of a workshop, this is biased towards confocal images and µCT (micro-computed tomography). The techniques are however applicable generally.
Getting Started with ImageJ / Fiji (PDF)
More techniques:
- Analysing matching regions of interest (PDF)
- Automatic alignment of images (PDF)
- Counting points in multiple regions of interest (PDF)
- Joining image arrays (PDF)
- Making a Look-Up-Table (PDF)
- Measuring area (PDF)
- Measuring area of a ringed selection (PDF)
- Measuring particle characteristics (PDF)
-
Quantifying 3D objects (PDF)
Explanation of 3D quantification functions (PDF) - Remove uncorrected background and montage (PDF)
- Showing colocalised regions (PDF)
- Spatial calibration of an image (PDF)
- Surface roughness (PDF)
- Thresholding (PDF)
Ilastik
![]() Ilastik is a simple, user-friendly tool for interactive image classification, segmentation and analysis. The program is undergoing quite rapid development, so check their website regularly.
Ilastik is a simple, user-friendly tool for interactive image classification, segmentation and analysis. The program is undergoing quite rapid development, so check their website regularly.
To get started with Ilastik we recommend visiting their website and we've selected some capabilities that may be of interest:
- Ilastik website
- Ilastic pixel classification is especially suited if the objects of interests are visually (brightness, color, texture) distinct from their surrounding. The algorithm is applicable for a wide range of segmentation problems that fulfil these properties.
- Ilastik carving is suited to greyscale images which do not exhibit clearly-delineated intensity zones, but have features separated by boundaries. TEM images, both 2D and 3D are good candidates for this method.
- Ilastik animal tracking digitally label animals in mazes and tracks and or quantifies where and when they go.
Osirix
 Osirix was designed for the medical profession, which means it's very good for viewing DICOM files and cataloging associated patient details in its database. It's also very useful for viewing many other forms of 3D data sets, whether from the µCT (micro-computed tomography) or confocal.
Osirix was designed for the medical profession, which means it's very good for viewing DICOM files and cataloging associated patient details in its database. It's also very useful for viewing many other forms of 3D data sets, whether from the µCT (micro-computed tomography) or confocal.
Free for research purposes, Osirix only runs on MacOS. It can be downloaded from Osirix's website in either 32 or 64 bit versions. The latter will cost you a few tens of dollars, but allows full use of your computer's RAM, which can be essential for large data sets.
Osirix technical sheet (Osirix website)
For some Windows alternatives, try looking at alternativeto.net/software/osirix
Exhibition images
See the images some of our talented staff produced:
2018 Science Festival photo exhibition
